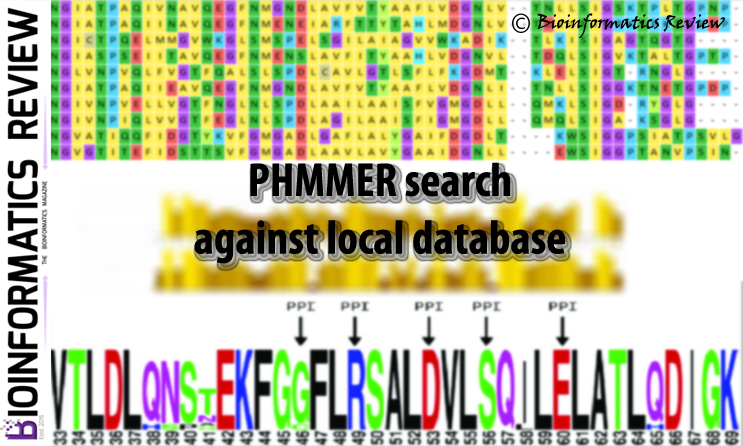













PHMMER is a sequence analysis tool used for protein sequences (http://hmmer.org; version 3.1 b2). It is available online as a web server and as well as...

The discovery studio (DS) visualizer [1] offers several features for analyzing docking results. In previous articles (“Tutorial: Vina Output Analysis Using PyMol” and “Video Tutorial: Autodock...

Reading a research article could be a problem for students starting their careers in research. Research papers are sometimes difficult to understand especially when you are...

Prottest3 is a software which is used to select a best-fit amino acid replacement model for a set of protein sequences [1]. ProtTest3 finds a best-fit...

Molecular docking is the most widely used technique which is used to predict the binding affinities and bound conformations of a ligand and protein complex. Autodock...

BLAST [1,2] is a local alignment tool widely used as a preliminary step for the identification of gene or protein functions. The command-line package of NCBI-Blast...

Cd-hit is one of the most widely used programs to cluster biological sequences [1]. It helps in removing the redundant sequences and provides better results in...

Here is a simple Perl script to search for motif patterns in a large FASTA file with multiple sequences.

The fast-emergence of predatory journals is a new problem in the scientific research area. These journals send attractive advertising emails to the authors to seek money...

A Ph.D. degree is considered difficult to get, you have to do a lot of hard work and give up certain things such as social life,...