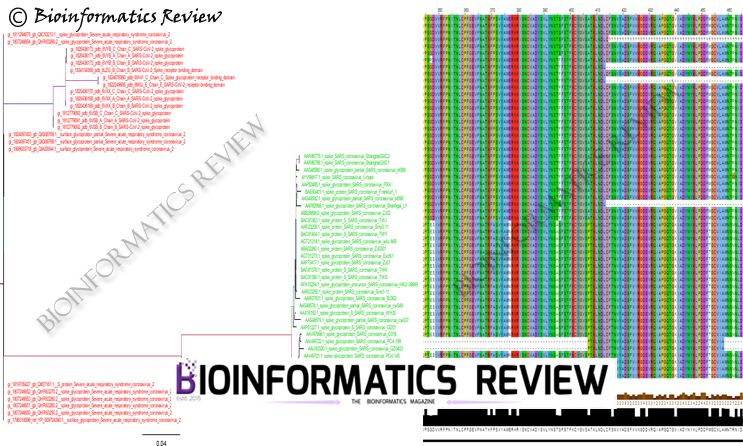
Protein sequence analyses include protein similarity, Protein function prediction, protein interactions, and so on. A new feature extraction model is developed for easy analysis of protein sequences.
This extraction model is known as FEGS (Feature Extraction based on Graphical and Statistical features) [1]. It represents protein sequences graphically based on their physicochemical properties and statistical features. By using these two properties/features, FEGS transforms a protein sequence into a 578-dimensional numerical vector.
How does FEGS work?
After taking protein sequences as input, FEGS starts building 158 space curves for the next protein sequence. After that, it builds L/L matrices and calculates normalized maximum eigenvalues. In the third step, it calculates the frequency of 20 amino acids and 400 dipeptides present in the sequence. It ultimately provides a frequency vector of that protein sequence. In the fourth step, it develops a feature vector of the sequence. It can later be subjected to phylogenetic analysis.
FEGS is a user-friendly software and freely downloadable on Sourceforge at https://sourceforge.net/projects/transcriptomeassembly/files/Feature%20Extraction/. FEGS’s performance has been tested on five different protein sequence datasets and it has shown the best performance amongst the other existing methods.
For more information, read here.
References
- Mu, Z., Yu, T., Liu, X. et al. (2021). FEGS: a novel feature extraction model for protein sequences and its applications. BMC Bioinformatics 22, 297.