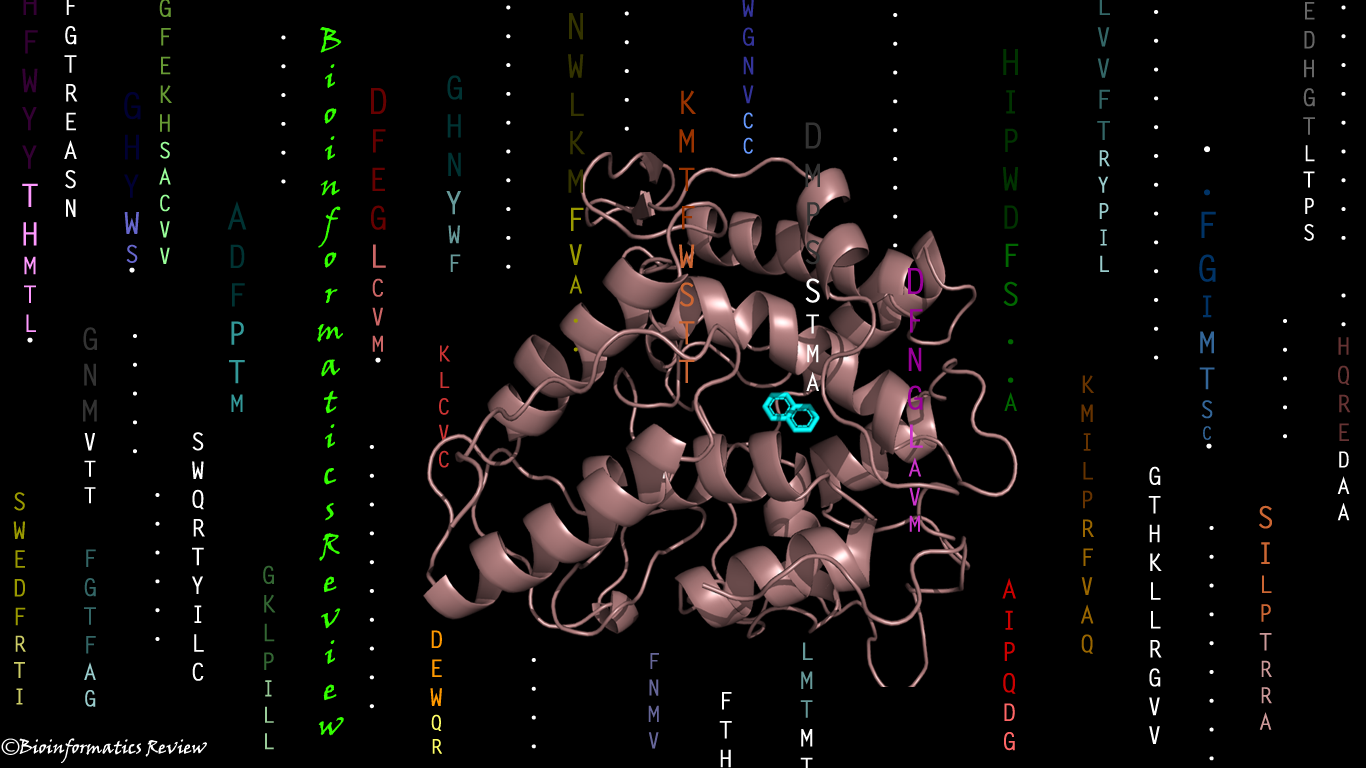
AI vs Physics in Molecular Docking: Towards Faster and More Accurate Pose Prediction
Abstract Molecular docking is a computational technique used in drug discovery to…
DockingAnalyzer.py: A New Python script to Identify Ligand Binding in Protein Pockets.
High-throughput virtual screening (HTVS) is a pivotal technique in drug discovery that…
CNN-DDI: A drug-drug interaction prediction method using convolutional neural networks
Drug-drug interaction (DDI) prediction is gaining much more interest in the drug…
DrugShot- A new web-based application to retrieve list of small molecules.
It is not an easy task to retrieve names of small molecules…
Most widely used tools for drug-drug interaction prediction
Drug-drug interaction is an important method in bioinformatics, especially in the case…
How to download small molecules from ZINC database for virtual screening?
It is difficult to manage thousands of compounds altogether while performing virtual…
Prepare receptor and ligand files for docking using Python scripts
Docking software such as Autodock4 and Autodock Vina require input receptor and…
Virtual Screening Methodology for Structure-based Drug Designing
Virtual High Throughput Screening (vHTS), also known as Virtual Screening (VS) is…
Beginner’s Guide for Docking using Autodock Vina
We have compiled all articles on docking into a Special Issue. This…
How to perform virtual screening using Autodock Vina?
Virtual screening is used to identify small molecules that are most likely…
Vina output analysis using Discovery Studio visualizer
The discovery studio (DS) visualizer offers several features for analyzing docking results.…
How to perform blind docking using AutoDock Vina?
Blind docking is done when the catalytic/binding residues are unknown in a…
Raccoon2: A GUI facilitating virtual screenings with Autodock and Autodock Vina
In the previous article, a new plugin called 'Raccoon' was mentioned, which…
How to install Raccoon plugin on Ubuntu for virtual screening using Autodock?
As mentioned in our previous articles, Autodock Vina is a very useful…
Video Tutorial: How to perform docking using Autodock Vina
This is a video added to our existing tutorial
Prediction of biochemical reactions catalyzed by enzymes in humans
There are many biological important enzymes which exist in the human body,…
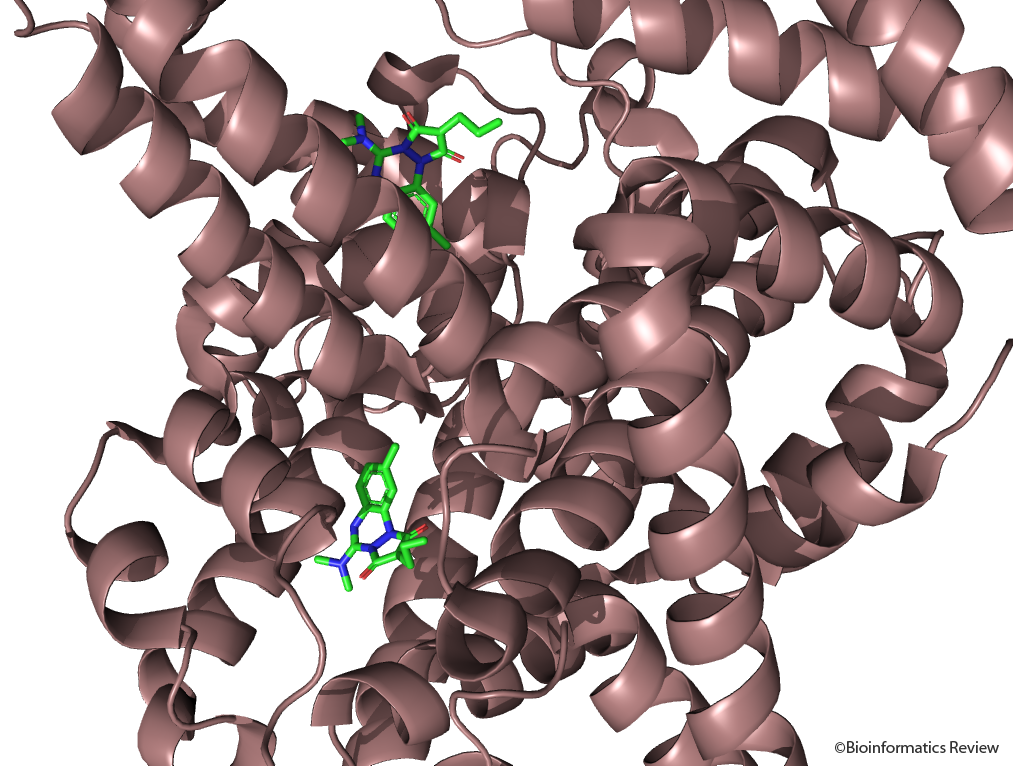
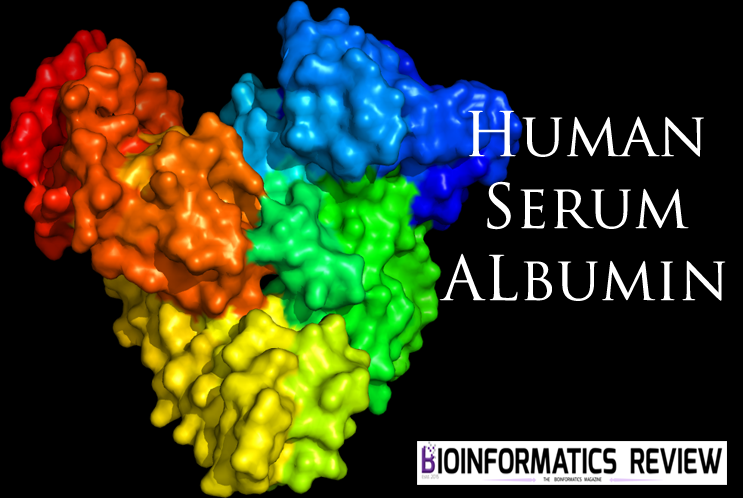
A review on the effects of CMPF binding with Human Serum Albumin
The interaction amongst plasma proteins and drugs is an imperative pharmacological parameter…
Systems pharmacology and drug development
Systems pharmacology is an emerging area in the field of medicinal chemistry…