Starting in Bioinformatics? Do This First!
I receive a lot of DMs asking for career guidance in the…
Video Tutorial: Fix broken Vina output in DS Visualizer.
This is a video tutorial on fixing the Vina docking output in…
How to Fix Broken Vina Docked Output in DS Visualizer: A Step-by-Step Guide
Molecular docking is a crucial method in computational drug discovery, allowing researchers…
CMake installation and upgrade: What worked & what didn’t?!
CMake is a widely used cross-platform build system that automates the process…
Common mistakes made during computational docking.
Computational docking is not a trivial task once we avoid making some…
How to download FASTA sequences from PDB for multiple structures?
In this article, we are going to download FASTA sequences for multiple…
How to install the LigAlign plugin on Pymol on Ubuntu (Linux)?
Few errors appear when we try to run the LigAlign plugin in…
How to make an impactful science presentation?
After your hard work, it is time to showcase your study and…
Basic bioinformatics concepts to learn for beginners
This article is for beginners who are stepping into the field of…
A Beginner’s Guide on How to Write Good Manuscripts
Drafting a manuscript could be a difficult task for beginners in the…
How to remove HETATMS and chains from PDB file?
This is a basic tutorial on removing the hetero-atoms (HETATMS) and chains…
Importance of Reasoning in Research
Research is considered complicated. As it involves reading multiple research articles and…
How to Find Binding Pocket/ Binding Site for Docking?
Finding binding sites/pockets in a target protein is one of the important…
Careers in Bioinformatics and Computational Biology
Bioinformatics is an interesting field of research combining biological sciences and computer…
Current Research Topics in Bioinformatics
Researchers working in the scientific area always want to explore new and…
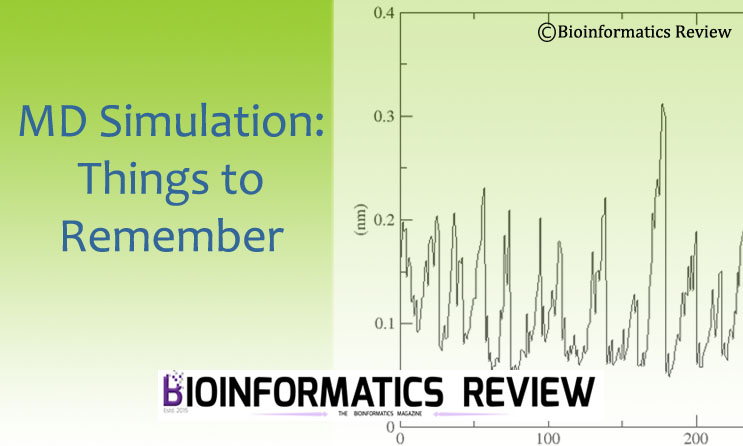
MD Simulation using GROMACS: Things to remember
Molecular dynamics (MD) simulation is considered amongst the important methods in bioinformatics.…
List of Bioinformatics Books for Beginners
It is difficult to decide which book to read to start learning…
Bioinformatics- Where & How to Start?
Bioinformatics being an interdisciplinary area of biological science and computer science may…
Common mistakes made during Autodock Vina Installation and Execution
Several errors occur while installing MGLTools and Autodock Vina on Ubuntu. We…
What does a bioinformatician do?
Bioinformaticians play an important part in data analysis and result interpretation in…