The plethora of research literature available to the modern day biologists provides the luxury to conduct a unique procedure- an analysis of the meta(data of data).
GWAS- Genome Wide Association Studies find their utility in aiding the researcher narrowing down to a specific biomolecule, to target for any curative or vague analytical procedure for any particular trait.
To make meta-analysis realistic and closer to truth one needs to scrutinize every individual study on some benchmark, VENICE CRITERIA here comes in handy.
Venice criteria can be understood as a set of three scores which are used to grade the evidence produced by the study. Each of these three score can attain a maximum of ‘A’ grade, followed by ‘B’, and ‘C’ based on how meticulous the study was.
- The first score is generated for “Amount”
- Second scoring is done for “Replication” and
- And final score is awarded for “Protection from bias”.
When trying to elaborate on each of these three grading criteria one must play in numerical quantities, and the details of the same follow
Amount
- ‘A’ grade is awarded for large scale evidence
1000 subjects, case: control= 1:1, for least common genetic group - For moderate evidence
100-1000 subjects, least common genetic group of interest - For little evidence
less than 100 subjects, least common genetic group of interest
Replication
- Extensively replicated study supported by at least 1 well conducted meta-analysis.
- Well conducted meta-analysis which may have faced some methodological limitations, or the studies have moderate inconsistency.
- The analyte lacks association or independently replicated study, has a flawed meta-analysis and no between study consistencies.
Protection from bias
Biases in studies creep in from researchers’ preconceived notions, and affect the compilation of data and declaration of result, much like previous two conditions a study must also be scrutinized for biases that may have crept in.
- Biases are minimized still can affect the magnitude, but probably not the presence of association.
- Based on the amount of missing information on generation of evidence, but the bias doesn’t clearly defer any associations.
- Evidence for bias is so heavy that it may affect the existence of any association between studies.
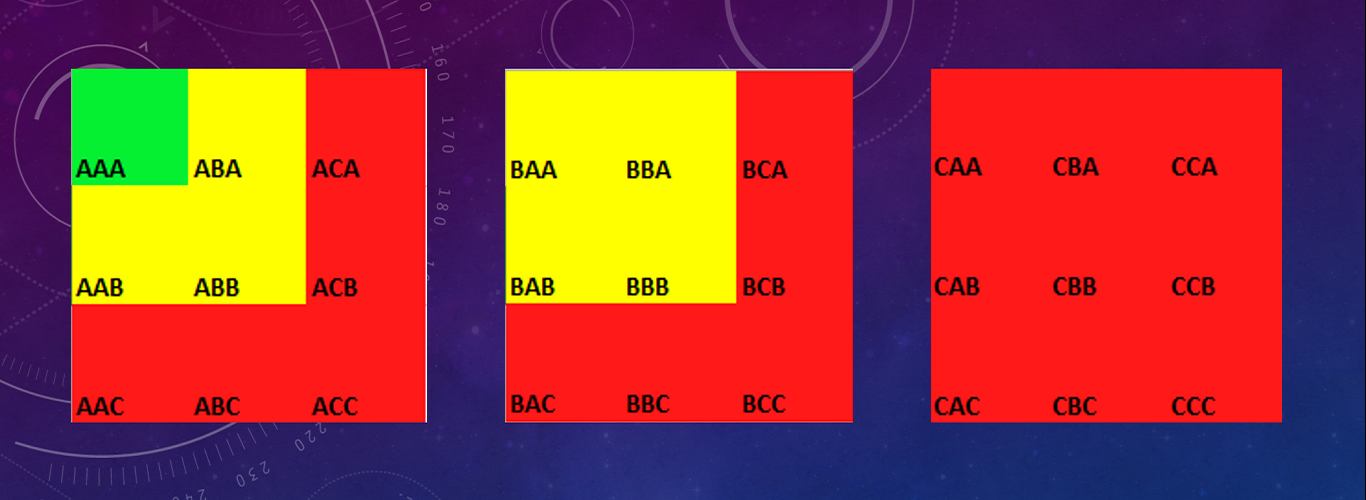
Thus the grades may be scored as follows-
AAA– strong evidence
AAB, ABA, ABB, BAA, BBA, BBB, BAB– moderate evidence
Rest all scores will be treated as poor, unreliable evidence.