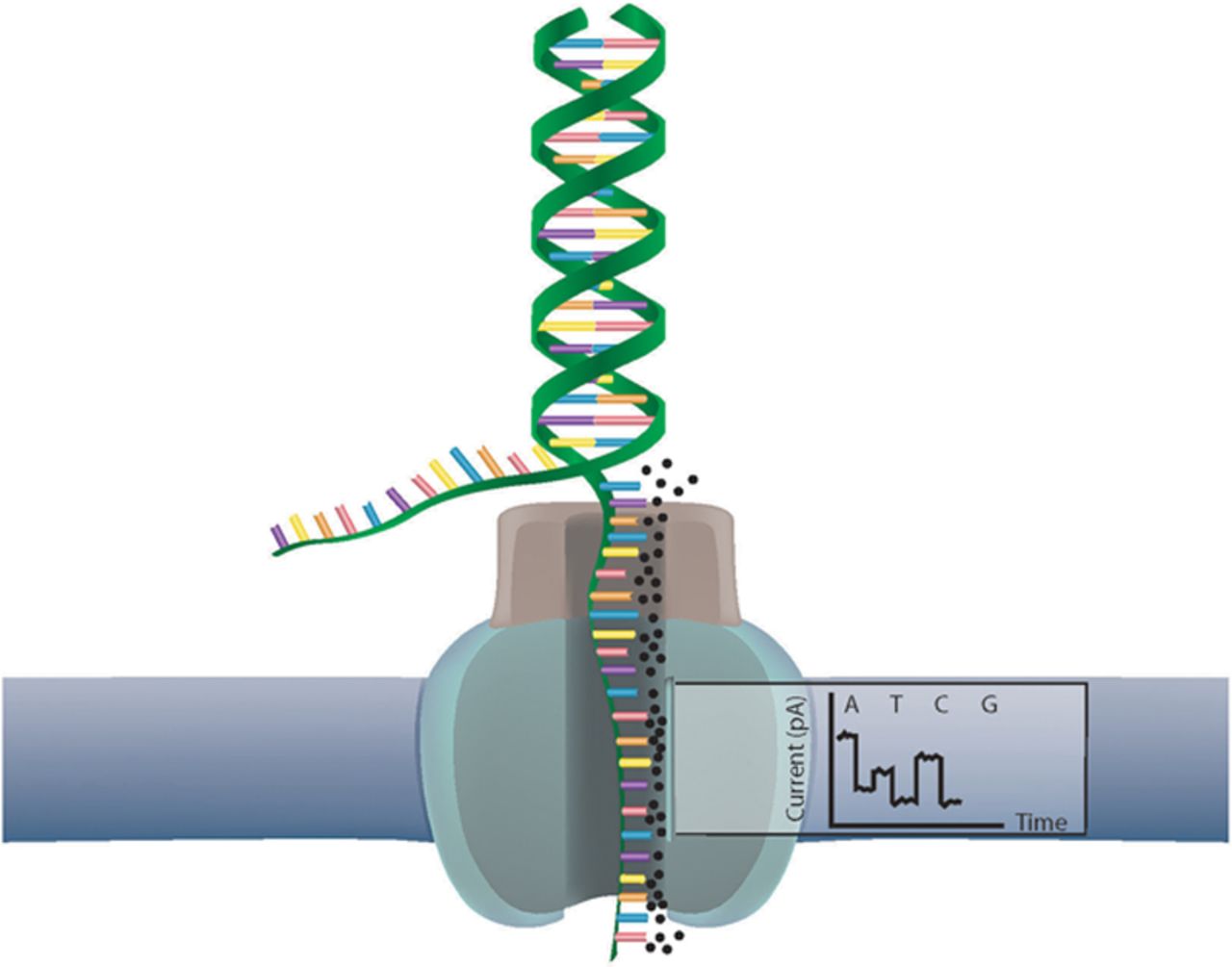
When it comes to bioinformatics algorithms, Genetic algorithms top the list of most used and talked about algorithms in bioinformatics. Understanding the Genetic algorithm is important not only because it helps you to reduce the computational time taken to get a result but also because it is inspired by how nature works.
In this article, you will learn how a genetic algorithm works, the basic concept behind it and we will also write a program to illustrate the concepts. You can skip the explanation if you already know the basic concepts of Genetic Algorithm
Genetic Algorithm was developed by John Holland. It uses the concepts of Natural Selection and Genetic Inheritance and tries to mimic the biological evolution. It falls under the category of algorithms known as Evolutionary Algorithms. It can be used to find a solution to the hard problems where we don’t know much about the search space.
Let us understand how a genetic algorithm works. For this, let us consider a cancer-associated gene expression matrix. This matrix contains all the known genes found in human being and their level of expression.
For a given problem, the genetic algorithm works by maintaining a set of candidate solutions and then applies three operators over them – Selection, Recombination, and Mutation, which are collectively known as the stochastic operators.
- Selection: In nature, if an organism is adapted to the environment, its population will grow relative to its quality of adaptation. This is referred to as selection. It means if a solution meets the conditional constraints, it is replicated at a rate which is proportional to the relative quality.
- Recombination: In nature, two similar chromosomes of the surviving individual exchange genes during sexual reproduction in a process known as Crossing Over. In GA we decompose two distinct solutions and randomly mix their parts to form novel solutions
- Mutation: Random changes in an existing chromosome may lead to some fitter individual. This concept is utilized to randomly perturbs a candidate solution
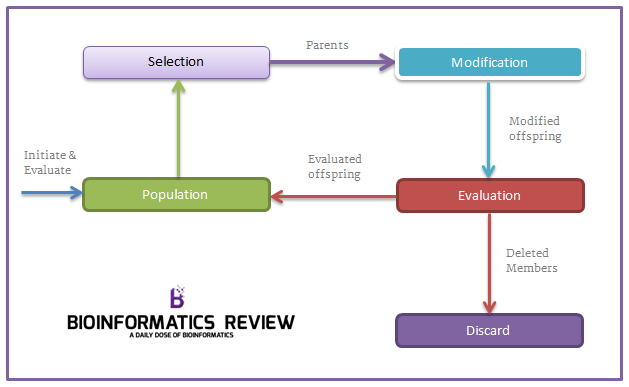
1. produce an initial population of individuals 2. evaluate the fitness of all individuals 3. while termination condition not met do 4. select fitter individuals for reproduction 5. recombine between individuals 6. mutate individuals 7. evaluate the fitness of the modified individuals 8. generate a new population 9. End while
Have a look at the Genetic Algorithm illustrated in the diagram below to understand it more clearly.
The program
We are going to implement the Genetic Algorithm and write a program in Perl for it. Although not purely applicable to a real-life problem, it should be sufficient to familiarize you with the Genetic Algorithm.
Suppose that you had a set of Gene expression data. The data is for all 25000 genes in the human genome and you want to find out what are the five values among all 25000 values whose sum can give you the highest number.
For the purpose of this program we will require four subroutines:
- Generate: It will generate chromosomes containing 5 values(specified in variable $GeneNumberConstraint) selected at random at positions
- Mutate: It mutates a chromosome at random position with a random value less than specified in $HighestMutationValue
- Survival Check: It checks if the newly formed chromosome is viable. i.e. It has a value that is up to a minimum specification. (Checking for fitness)
- Recombine: It will form new combinations from the existing chromosome by crossing them over with each other.
The Code
If you wish, you can download the Perl code on GitHub https://github.com/bioinformaticsreview
/geneticalgorithm
So here is the final code implementing Genetic Algorithm in Perl:
# Written by Tariq Abdullah
# Author, Bioinformatics Review
# tariq@bioinformaticsreview.com
# www.bioinformaticsreview.com
$CurrentHighest=0;
@GeneExpressionData = (1,3,8,5,2,4,46,6,78,7,9,9
,0,1,1,1,5,59,9,97,7,6,5,45
,4,3,23,2,22,2,2,4,5,5,6,54);
@SolutionSpace = ();
$HighestMutationValue = 110;
$GeneNumberConstraint = 5;
$InitialThreshold = 10;
$genes = scalar @GeneExpressionData;
@chromosome = ();
$sum = 0;
$steps= 10;
print "The Total Genes are: $genes\n";
generate();
$steps = 10;
for($p=0;$p<=$steps;$p++){
generate();
SurvivalCheck();
mutate();
SurvivalCheck();
recombine();
SurvivalCheck();
}
print "\n\n Genetic Algorithm Result
\n\n\n\t\tHighest Detected: $CurrentHighest in $steps Steps\n\n";
sub mutate{
$randpos = int(rand($gene));
$n = int(rand($HighestMutationValue));
$chromosome[$randpos] = $n;
print "\n Mutation Took Place in Chromosome @chromosome ";
}
sub recombine
{
print "\nRecombining\n\n";
@chromosome1 = $SolutionSpace[int rand($p)];
@chromosome2 = $SolutionSpace[int rand($p)];
print "Random Sequence Chromosome from Solution Space: @chromosome1 and @chromosome2";
for($i=0; $i<=$GeneNumberConstraint/2; $i++){ my $random_number = int(rand(3)) + 1; $pos1 = int(rand($GeneNumberConstraint)); $pos1 = int(rand($GeneNumberConstraint)); $swap = $chromosome1[$pos1]; $chromosome1[$pos1] = $chromosome2[$pos2]; $chromosome2[$pos2] = $swap; } print "The Recombination led to @chromosome"; @chromosome = (); @chromosome = @chromosome1; } sub SurvivalCheck{ $sum = 0; foreach $val (@chromosome){ $sum += $val; } if($sum>$CurrentHighest){
$CurrentHighest = $sum;
push @SolutionSpace, @chromosome;
print "\nIndividual is alive! \nCurrent Highest Expression: $CurrentHighest"; return 1;
}
else{
print "\nSpecies Didn't Survive! \n";
return 0;
}
}
sub generate{
@chromosome = ();
for($i=1;$i<=$GeneNumberConstraint;$i++){
$n = int(rand($genes));
push @chromosome, $GeneExpressionData[$n];
$sum += $GeneExpressionData[$n];
}
print "\n\n\nGenerated Chromosome: @chromosome \n";
}
That’s all! Feel free to comment and discuss if you have any confusion. Like this article? Share it.. ha?