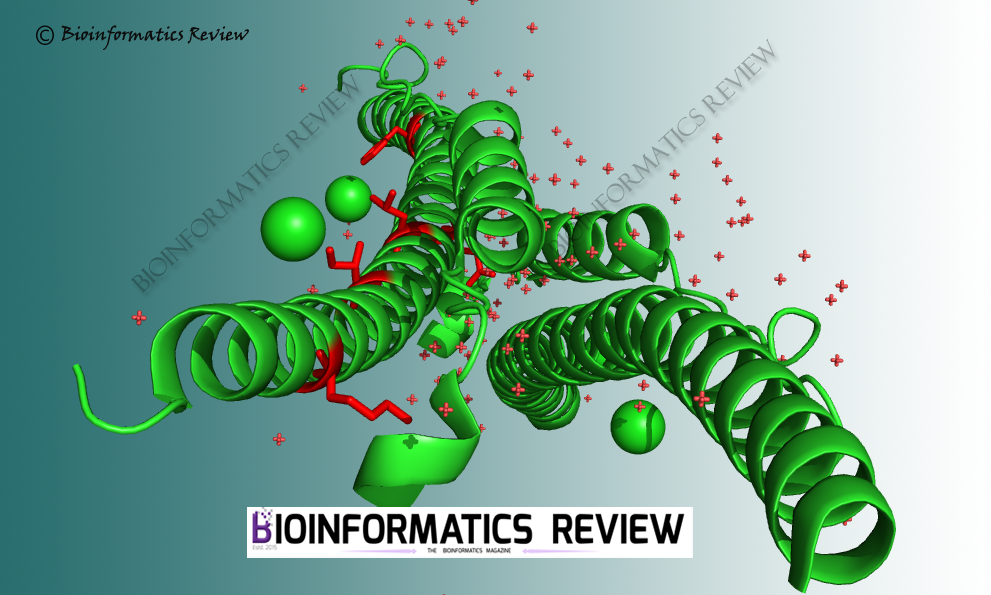
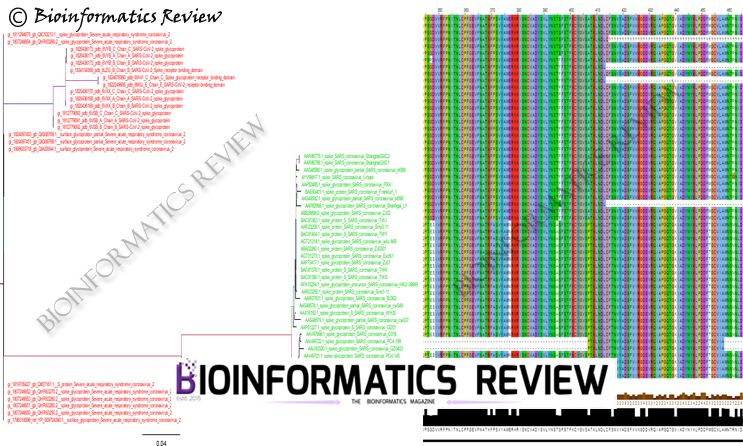
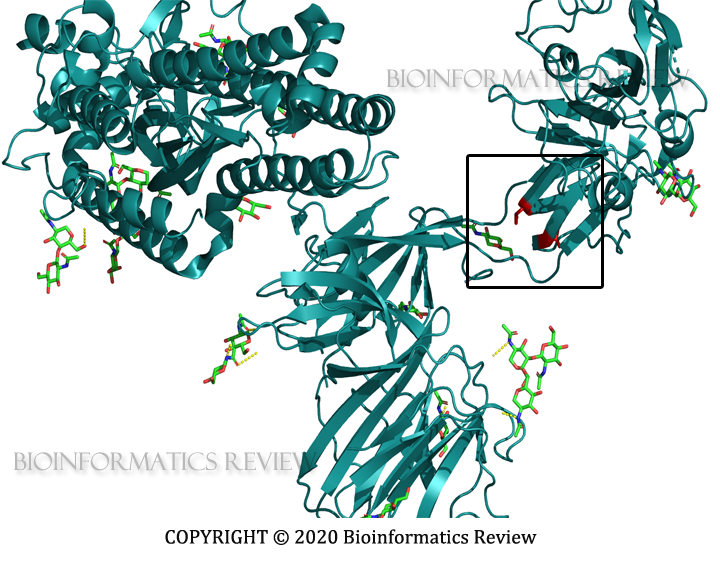
In our last article, we mentioned the selection pressure analysis of SARS-CoV-2 spike glycoproteins. Now, we have analyzed spike glycoprotein sequences of SARS-CoV. No single purifying selection site was found in SARS-CoV spike glycoproteins as was revealed from our last analysis of SARS-CoV-2 spike glycoproteins.
The analysis was performed using Hyphy [1] on the Datamonkey server [2]. BUSTED [3], MEME [4], and FUBAR [5] methods were used. MEME [4] and FUBAR [5] found 6 and 2 sites respectively under positive selection. The threshold was set to 0.05 (p-value) for MEME and 0.9 (posterior probability) for FUBAR. Additionally, unlike SARS-CoV-2, BUSTED [3] found evidence of selection on the tested branches of the phylogeny of SARS-CoV spike glycoproteins. It implies that there is at least one site on one branch that has experienced gene-wide episodic positive selection.
Interestingly, negative selection has not been found in both SARS-CoV and SARS-CoV-2 spike glycoproteins. Besides, only a small number of sites have been found to be experienced positive selection. These sites of SARS-CoV spike proteins under selection are being analyzed further.
References
- Pond, S. L. K., & Muse, S. V. (2005). HyPhy: hypothesis testing using phylogenies. In Statistical methods in molecular evolution (pp. 125-181). Springer, New York, NY.
- Weaver, S., Shank, S. D., Spielman, S. J., Li, M., Muse, S. V., & Kosakovsky Pond, S. L. (2018). Datamonkey 2.0: a modern web application for characterizing selective and other evolutionary processes. Molecular biology and evolution, 35(3), 773-777.
- Murrell, B., Weaver, S., Smith, M. D., Wertheim, J. O., Murrell, S., Aylward, A., … & Scheffler, K. (2015). Gene-wide identification of episodic selection. Molecular biology and evolution, 32(5), 1365-1371.
- Murrell, B., Wertheim, J. O., Moola, S., Weighill, T., Scheffler, K., & Pond, S. L. K. (2012). Detecting individual sites subject to episodic diversifying selection. PLoS genetics, 8(7).
- Murrell, B., Moola, S., Mabona, A., Weighill, T., Sheward, D., Kosakovsky Pond, S. L., & Scheffler, K. (2013). FUBAR: a fast, unconstrained bayesian approximation for inferring selection. Molecular biology and evolution, 30(5), 1196-1205.