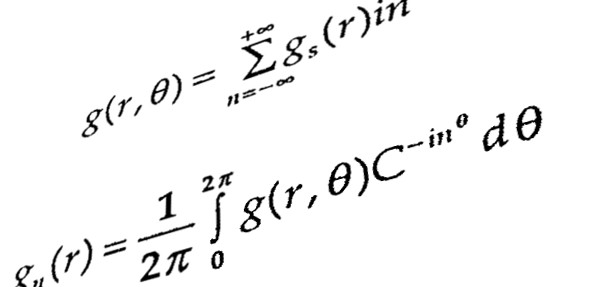Geneious: A platform for Comprehensive Analysis and Organization of Genes
The two main functions of bioinformatics are the organization and analysis of…
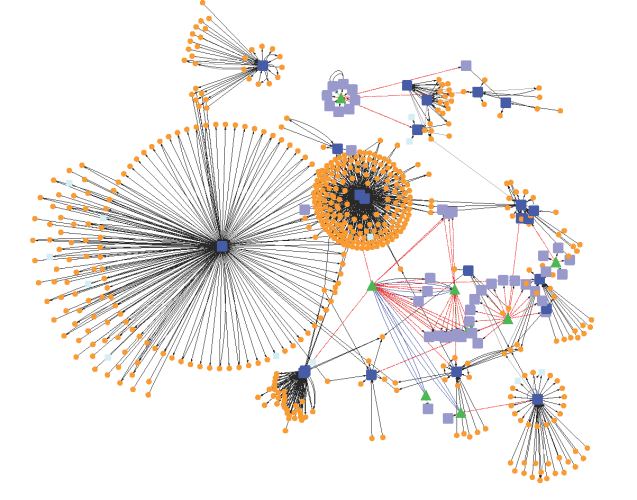
MOTIF: Functional Unit of An Interaction Network
In a network, integration of elements and interacting components enables identification of…
Editorial, Looking forward and beyond
It’s all warm in BiR as we are inching towards next-to-next big…
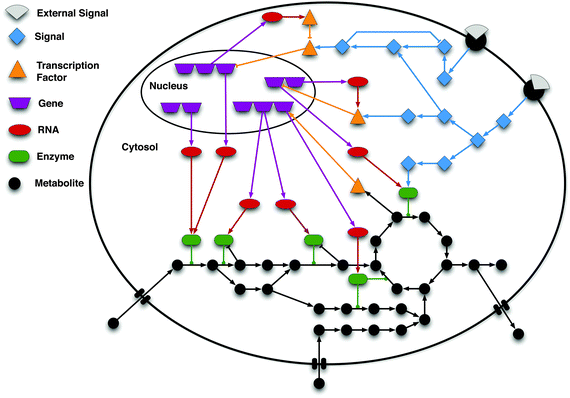
Network Biology: Get Together of Macromolecules
A network is a group of two more than two interacting components.…
Molecular Evolutionary Genetic Analysis
MEGA: Molecular Evolutionary Genetic Analysis It is important to know the basic…
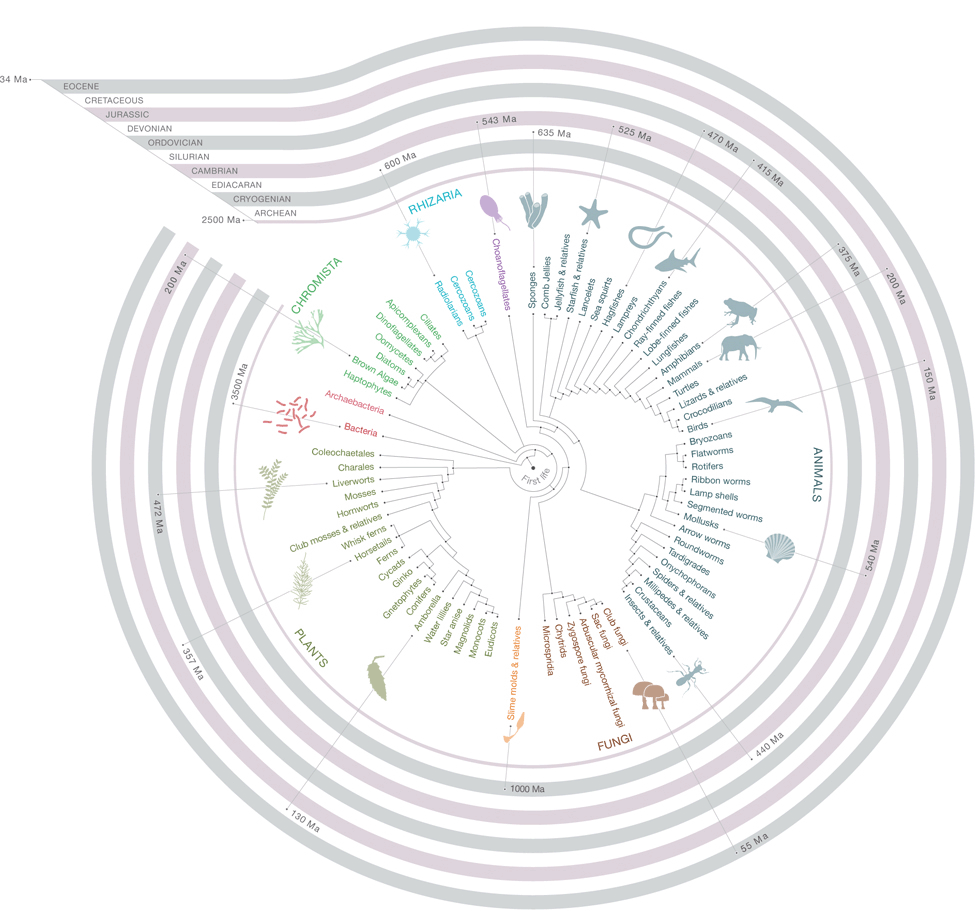
Monogenea: An Easy Portfolio for Molecular Phylogenetics
Estimation of present day diversity of organism and understanding their diversity forms…
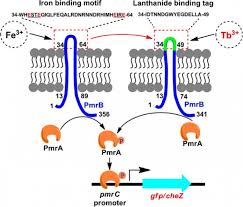
Two Components System: Potential Drug Target in Mycobacterium tuberculosis
The genomic complexity and unknown functions of proteins/genes in Mycobacterium tuberculosis (Mt) has…
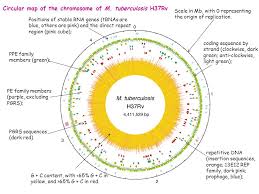
Explore Tuberculosis: A Systems Biology Approach
Systems biology is not sufficient to full fill the requirement of molecular…
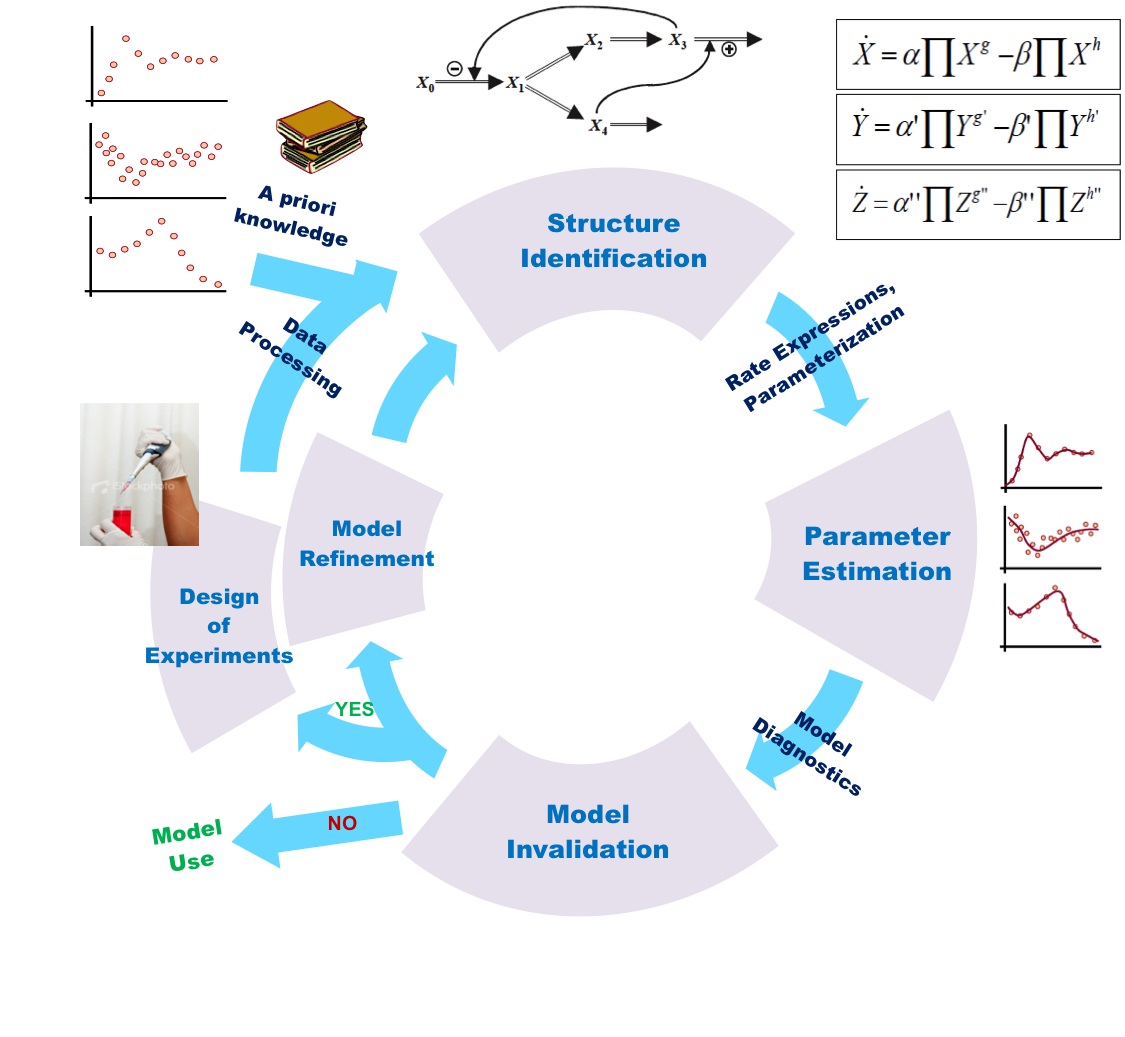
Introduction to Mathematical Modelling (Last Part)
In the previous section, mathematical modeling was exemplified by metabolic process and…
Introduction to Mathematical Modelling Part-3
Derivation of Mathematical Equations for Understanding Systems Behaviour: Depending upon the nature…
Introduction to mathematical modelling – Part 2
Gathering of Dynamic/Kinetic information In the previous section you might have noticed…
Basics of Mathematical Modelling – Part 1
Biochemical processes are simply complex, and their apparent feature does not easily…