Alternative Splicing plays a very essential role in proper functioning of eukaryotic cells. It acts as a regulatory mechanism for gene expression and any kind of disruption in this mechanism may lead to human diseases. Alternative splicing of pre-mRNA is a major source of genetic variation in human beings and disruption of the splicing process may cause human diseases such as cancer. Cancer-associated variation which may occur at different levels of gene regulation, particularly during the processing of pre-mRNA into mature mRNAs. So, better understanding of these mechanisms may provide insights into disease causes and development.
TIN is a new R package which enables to analyze TIN from the expression data. TIN is a software package of R modules that uses a framework to analyze expression level data.
WORKFLOW:
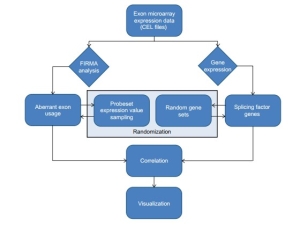
TIN uses raw expression data (cell intensity,CEL files) as input and applies the FIRMA method (i.e., a method for detection of alternative splicing) estimating the expression levels of transcriptome and the alternative splicing patterns between samples. FIRMA method gives a FIRMA score to each exon sample combination, which is based on the deviation of probes systems from the expected gene expression level. Thus, FIRMA score is the relative ratio between exon expression level and corresponding gene expression level. If FIRMA shows a strong positive score, then the differential exon is included and if it shows a negative score, then it implies that exon is skipped.
Since alternative splicing is mediated by several splicing factors and proteins which remove introns from the pre-mRNA then joining the exons of mRNA together. Therefore, TIN basically test the association between splicing factor expression levels and amount of abnormal exon usage among the samples. For this, correlation between abnormal exon usage amounts and splicing factor expression levels tested across all samples is calculated. If the correlation is considerably lower, it indicates that the aberrant amounts of exon expression may be due to splicing factor expression. After that, correlation is tested by using random gene sets, if the correlation is poor then it gives an indication that the abnormal exon usage can be attributed to the expression levels of the splicing factor genes.
Fig.1 Workflow of TIN
This is how by analyzing the gene expression levels and alternative splicing patterns we can easily monitor a developing disease or it can be predicted at an very early stage.
For further reading, click here.
Note:
An exhaustive list of references for this article is available with the author and is available on personal request, for more details write to muniba@bioinformaticsreview.com